Take our quiz to test your knowledge about Covid-19 and SARS-CoV-2 from Here
Methods for Testing SARS-CoV-2 for Covid-19 Diagnosis
There are 2 main kinds of tests for SARS-CoV-2. One type involves detection of the virus itself (viral RNA or antigen) and the other type involves detection of the human immune response to infection (antibodies or other biomarkers).
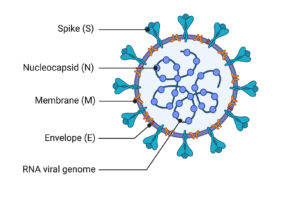
Standard confirmation of acute SARS-CoV-2 infections is based on the detection of unique sequences of virus RNA by nucleic acid amplification tests (NAATs), such as reverse-transcription polymerase chain reaction (RT-PCR). The viral genes targeted so far include the N, E, S and RdRP genes.
Reverse-Transcription Polymerase Chain Reaction (RT-PCR)
Polymerase chain reaction (PCR) is a molecular technique which allows production of million copies of a specific DNA sequence from initially smallest sample, within few hours.
| Abbreviation | Full Name |
|---|---|
| PCR | Polymerase Chain Reaction |
| qPCR | Quantitative Polymerase Chain Reaction |
| rPCR | Real-time Polymerase Chain Reaction |
| RT-PCR | Reverse transcription Polymerase Chain Reaction |
| rRT- PCR | Real-time Reverse transcription Polymerase Chain Reaction |
| RT-qPCR | Reverse transcription-quantitative Polymerase Chain Reaction |
Reverse transcription polymerase chain reaction (RT-PCR) is a laboratory technique combining reverse transcription of RNA into DNA (called complementary DNA or cDNA) and amplification of specific DNA targets using polymerase chain reaction (PCR). This is achieved by monitoring the amplification reaction using fluorescence, a technique called real-time PCR or quantitative PCR (qPCR).
Principle of RT-PCR
The PCR involves the primer mediated enzymatic amplification of DNA. PCR is based on using the ability of DNA polymerase to synthesize new strand of DNA complementary to the offered template strand.
Reverse transcription PCR (RT-PCR) is used when the starting material is RNA. As viruses like SARS-CoV-2 contain RNA as their genetic material, RT-PCR is used. In this method, A sample is collected from the parts of the body where the COVID-19 virus gathers, such as nasopharyngeal or oropharyngeal swab. RNA is extracted removing undesired components by chemical treatment. This extracted RNA is a mix of the person’s own genetic material and, if present, the virus’s RNA. RNA is first transcribed into complementary DNA (cDNA) by reverse transcriptase from total RNA or messenger RNA (mRNA). The cDNA is then used as the template for the PCR reaction.
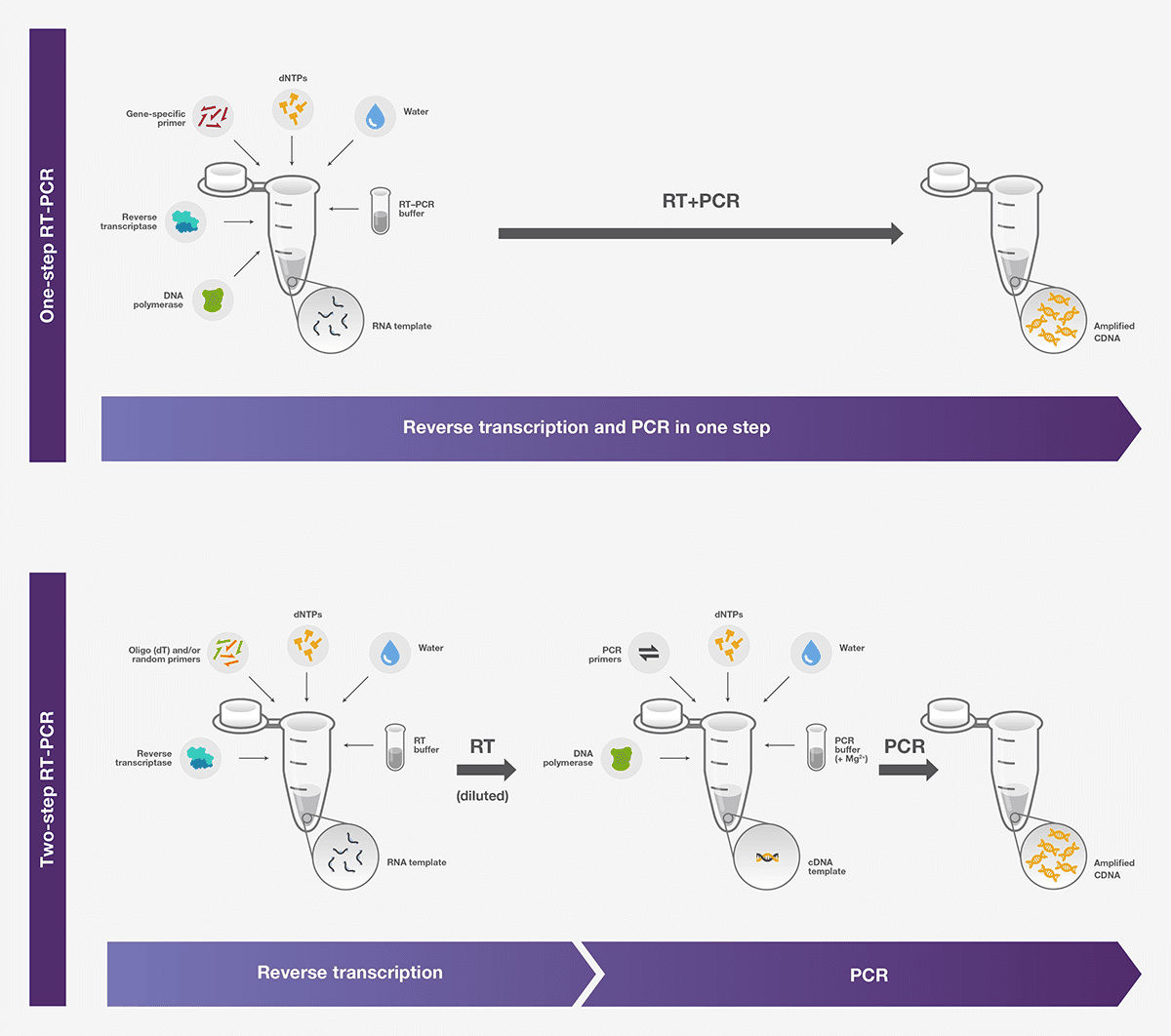
RT-PCR can be performed by two methods: one-step or a two-step assay. One-step assays combine reverse transcription and PCR in a single tube and buffer, using a reverse transcriptase along with a DNA polymerase. In two-step assays, the reverse transcription and PCR steps are performed in separate tubes, with different optimized buffers, reaction conditions, and priming strategies.
Components of RT-PCR
The RT-PCR reaction requires the following components:
-
RNA Template
The single stranded RNA (ssDNA) of interest, separated from the sample.
-
Primers
A primer is a short nucleic acid sequence that provides a starting point for DNA synthesis. It helps to provide 3′-OH group to add the first nucleotide. Primer for reverse transcription anneals to the template mRNA strand and provide reverse transcriptase enzymes a starting point for synthesis. These are complementary to the 3’ ends of the sense and anti-sense strands of the target sequence.
-
Reverse Transcriptase
Reverse transcriptase is an RNA-dependent DNA polymerase, catalyzing DNA synthesis using RNA as the template.
-
DNA Polymerase
Usually a thermostable Taq polymerase that can function at a temperature optimum of about 70°C and does not rapidly denature at high temperatures (98° C).
-
Deoxynucleotide triphosphates
Single units of the bases A, T, G, and C (dATP, dTTP, dGTP, dCTP) provide the energy for polymerization and the building blocks for DNA synthesis.
-
Buffer system
Includes magnesium and potassium to provide the optimal conditions for DNA denaturation and renaturation; also important for polymerase activity, stability and fidelity.
-
Thermocycler
A thermal cycler (also known as a PCR machine or thermocycler) is a laboratory instrument that heats and cools samples in repetitive cycles to facilitate DNA or RNA amplification through the polymerase chain reaction.
Stages of RT-PCR
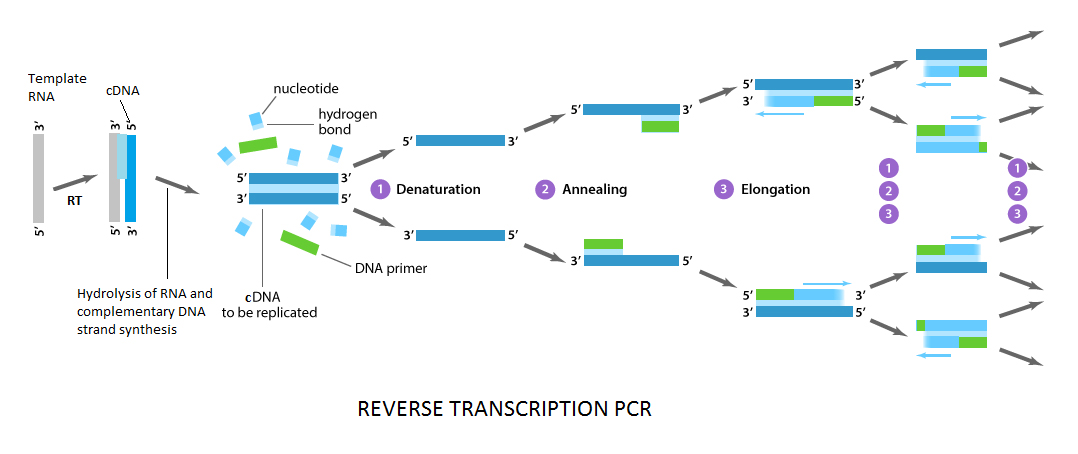
The process of the PCR is subdivided into three stages: denaturation, annealing and elongation. RT–PCR is a variation of PCR which use the same process except that RT–PCR has an added step of reverse transcription of RNA to DNA at first to allow for amplification. The first cycle is reverse transcription to synthesize cDNA. The second cycle is initial denaturation. During this cycle reverse transcriptase is inactivated. The next 40 to 50 cycles are the amplification program, which consists of three steps: (1) denaturation, (2) annealing, (3) elongation.
-
Reverse Transcription:
Reverse transcriptase enzyme synthesizes a complementary DNA (cDNA) strand with nucleotides, extending from the primer. It occurs between 40°C and 50°C, depending on the properties of the reverse transcriptase enzyme utilized. This is one time reaction and the product is mRNA:cDNA hybrid. After the reverse transcription, the mRNAs are hydrolyzed and single-stranded cDNAs are then replicated by the DNA polymerase during a first temperature cycle.
-
Denaturation:
All the component mixture is heated to 94°C for 15-30 seconds. This causes breakdown of hydrogen bonds and double stranded cDNA is denatured to single strands. These will act as templates for the production of the new strands of DNA.
-
Annealing:
The reaction temperature is rapidly lowered to 54-60°C for 20-40 seconds. Primers bind to the target DNA sequences and initiate polymerization.
-
Elongation/Extension:
This step usually occurs at 72-80°C (most commonly 72°C). The DNA Taq polymerase enzyme sequentially adds bases to the 3′ end of primer, extending the DNA sequence in the 5′ to 3′ direction. DNA polymerase will add about 1,000 bp/minute under optimal conditions.
The result of one cycle of PCR is two double-stranded sequences of target DNA, each containing one newly made strand and one original strand. Upon denaturation, these new fragments also serve as templates, Each cycle doubles the previous number.As the cycles are repeated, more and more copies are generated and the number of copies of the template is increased exponentially.
Detection in Real-Time RT-PCR (rRT-PCR/RT-qPCR)
Real Time Real-time reverse-transcription PCR (rRT-PCR) is the technique of collecting data throughout the PCR process as it occurs, thus combining amplification and detection into a single step. This is using fluorescent dyes that yield increasing fluorescent signal in direct proportion to the number of PCR product generated. Fluorescent reporters used in real-time PCR include double-stranded DNA (dsDNA)- binding dyes, or dye molecules attached to PCR primers or probes that hybridize with PCR product during amplification. This value is usually referred to as cycle threshold (Ct), the time at which fluorescence intensity is greater than background fluorescence. Greater the quantity of target DNA in the sample, there will be significant increase in fluorescent signals earlier, yielding a lower Ct.

Be the first to comment